In my research, I explore how microbial dynamics may be used to better predict the fate of contaminants in aquatic ecosystems.
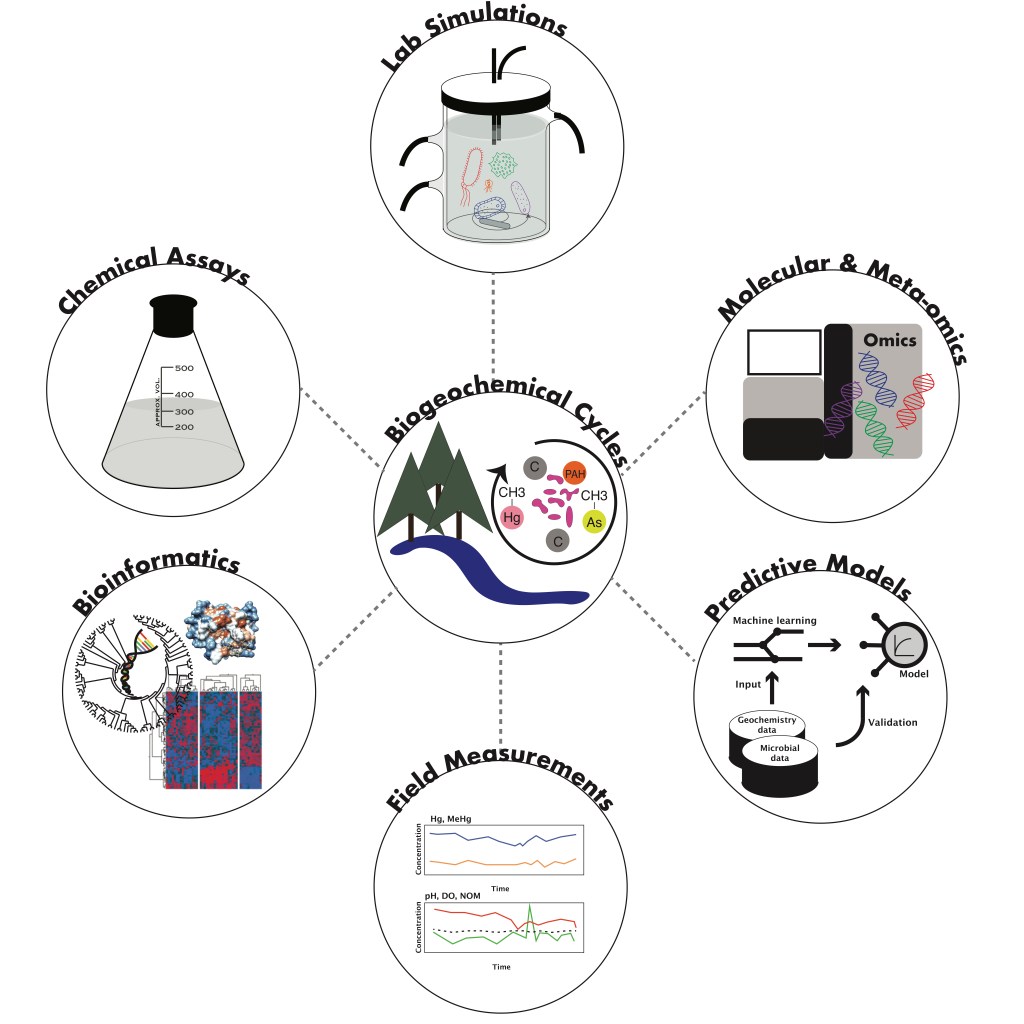
Microbes play a significant role in controlling the form and fate of metals in the environment. In my research, I utilize various molecular and meta-omic methods for linking microbial community dynamics to metal biogeochemistry in aquatic ecosystems. My recent research is primarily focused on studying the microbial genetics of mercury methylation. Methylmercury is a highly toxic organic form of mercury that accumulates in the food web, including rice and fish, posing a threat to human and ecosystem health. I am interested in understanding why microbes produce this more toxic form of mercury and how we may limit its production in the environment. I often use a combination of amplicon sequencing, meta-omics analyses, microcosm manipulation experiments, in-depth omics analyses, and empirical modelling to investigate how microbes cycle mercury. These methods enable the identification and quantification of mercury-methylating microbial groups, the assessment of genetic potential and activity of the microbial community, the study of how environmental factors impact microbial dynamics, and the development of predictive models for net methylmercury accumulation. By analysing the composition and activity of microbial communities, and their genetic potential for mercury transformations, my research contributes to our understanding of the environmental drivers of methylmercury production and its implications for ecosystem health. The biogeochemical mercury cycle significantly overlaps with cycling of other elements, including arsenic, iron, sulfur, nitrogen, and carbon. Therefore, outcomes from my research reach beyond mercury, contributing to our fundamental knowledge of microbial processes in natural environments and enabling more robust and accurate predictive models that can be applied at the ecosystem and global scale.
Potential Student Projects
I can supervise Honours, Masters, and PhD research projects focused on linking microbial genetics to ecosystem health. I am especially interested in projects that investigate the role microorganisms play in regulating biogeochemical cycles in aquatic ecosystems, including how microbial transformations of emerging contaminants and toxic metals affect their accumulation in ecosystems and food webs. My current research is focused on microbial transformations of mercury and arsenic in freshwater and marine ecosystems. However, I am capable of supervising student research projects in broader areas of microbial ecology or biogeochemistry. My broader interests span an array of topics in environmental microbiology, including how microbiome structure contributes to ecosystem functioning and understanding the response of microbial communities to climate change or disturbance. I can also supervise projects that have a more applied focus, for example bioremediation of acid-mine drainage or coal-ash spills.
Please get in touch by email if you are interested in discussing potential student research projects.
